Manas Santra, Ph.D.
manas@nccs.res.in
Research Areas
Biology of Cancer and chronic Diseases, Genome Architecture and Regulation
Education and Experience
2021 to present: Scientist F, National Centre for Cell Science, Pune, India.
2015 to 2020: Scientist E, National Centre for Cell Science, Pune, India.
2010 - 2015: Scientist D, National Centre for Cell Science, Pune, India.
2006 - 2010: Postdoctoral Fellow, University of Massachusetts Medical School, USA
2001 - 2006: Ph. D., Indian Institute of Technology Bombay, Maharashtra
1999 - 2001: M. Tech. (2001), Indian Institute of Technology, Maharashtra
1996 - 1998: M. Sc. (Chemistry), Vidyasagar University, West Bengal
1993-1996: B. Sc. (Chemistry), Vidyasagar University, West Bengal
Research
A. Gene Regulation by RING Finger E3 Ubiquitin Ligase (SCF and APC/C):
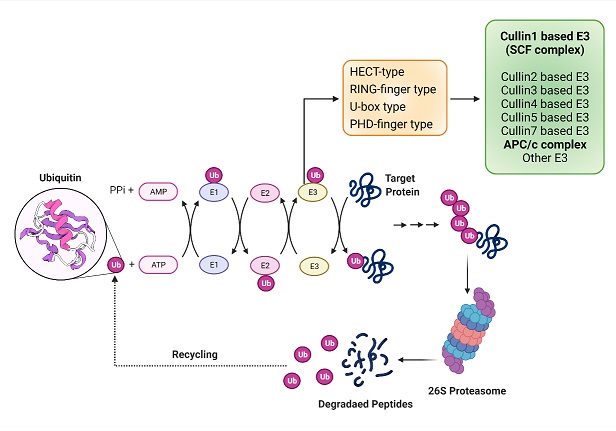
Ubiquitin ligases are specific class of proteins mainly contributing to the proteasomal degradation of many key proteins involved in the cell cycle regulation as well as many signalling pathways. These are well suited to regulate molecular networks that operate at post-translational level. SCF (Skp1-Cul1-F-box) and APC/C ubiquitin ligase complexes are best studied E3 ubiquitin ligase. The F-box proteins act as a substrate receptor and determine the substrate specificity for the complex. Deregulation of this proteolytic system might result in uncontrolled proliferation, genomic instability and diseases like cancer. Although there are 69 F-box genes present in the human genome, majority of them remain uncharacterized. The inherent interacting capability of these proteins makes them important regulators of various important cellular processes like cell cycle progression, apoptosis, senescence, metabolism etc. Our lab is interested in identifying the F-box proteins having potential role in tumor biology and elucidating their molecular mechanism through different molecular cell biology, biochemistry and proteomic approaches.
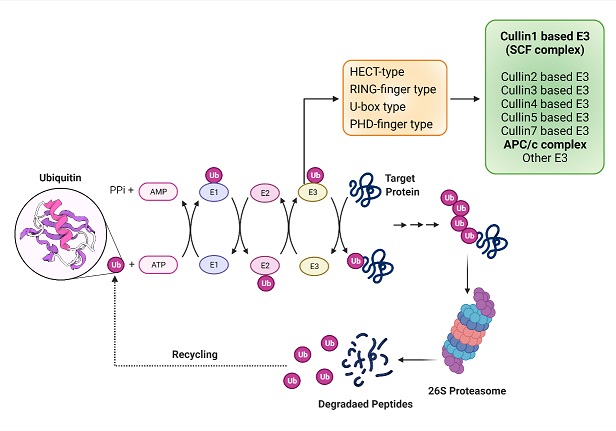
Schematic model for ubiquitination of proteins by E3 ligases
Schematic of SCF (SKP1-Cullin1-F-box) complex mediated protein degardation
B. Understanding the Role of Epigenetic Gene Regulation in Cancer:
Epigenetics is the study of heritable changes in gene expression without altering the DNA sequence. Along with the genetic, epigenetic mechanism of gene regulation plays an important role in tissue specific spatio-temporal gene expression pattern, perturbation of which leads to diseases like cancer. Cancer arises due to the inactivation of the tumor suppressor genes and activation of oncogenes. In many instances, loss of function of tumor suppressor genes is due to epigenetic alterations. Our lab is interested in deciphering how and why the tumor suppressor genes are epigenetically silenced in cancer and their consequences.

Epigenetic reprogramming during cancer development
C. Identification and elucidation of the role of Protein Phosphatase 1 regulatory subunits in Cancer:
Proteins are the functional unit of the cell and post translational modifications (PTMs) dictate their cellular functions. Among the different PTMs, phosphorylation (mediated by kinases) and dephosohorylation (mediated by phosphatases) are the major modifications. The phosphatases are key players in maintaining normal cellular homeostasis, as illustrated by the dephosphorylation of p53 to bring down the cell to normal steady state after DNA damage response. There are many protein phosphatases (PP) in the cell, namely PP1, PP2, PP4, PP5, PP6 etc. Protein phosphatase 1 (PP1) belongs to the family of serine-threonine phosphatases and forms a major fraction of the cellular phosphatase pool. It plays significant role in almost all important cellular functions such as cell cycle regulation, stress response and many more. Differential recruitment of regulatory subunits of PP1 dictates the substrate specificity for its function. Taking into consideration the significance of PP1 in cellular functions, our lab is interested in exploring the repertoire of PP1 regulatory subunits having a role in cancer biology.

Schematic of dephosphorylation of substrate protein by protein phosphatase 1 (PP1)
Publications
|
1. Islam S, Dutta P, Sahay O, Gopalakrishnan K, Roy Muhury S, Parameshwar P, Shetty P, Santra MK. (2022) Feedback-regulated transcriptional repression of FBXO31 by c-Myc triggers ovarian cancer tumorigenesis. Int J Cancer. 150, 1512 – 1524. |
|
2. Agrawal, Y., Nadkarni, K. S., Gupta, N. A., Manne, R. K. and Santra, M. K. (2022) F-box protein FBXO41 plays vital role in arsenic trioxide-mediated autophagic death of cancer cells. Toxicol Appl Pharmacol. 2022 Apr 15;441:115973. |
|
3. Sahay, O., Barik, G. K., Sharma, T., Pillai, A. D., Rapole, S. and Santra, M. K. (2022). Damsel in distress calling on her knights: Illuminating the pioneering role of E3 ubiquitin ligases in guarding the genome integrity. DNA Repair (Amst). 2022 |
|
4. Islam, S., Dutta, P., Sahay, O., Santra M. K. (2021) β-TrCP1 facilitates cell cycle checkpoint activation, DNA repair and cell survival through ablation of β-TrCP2 in response to genotoxic stress. J. Biol. Chem. 296, 100511. |
|
5. Manne RK, Agrawal Y, Malonia SK, Banday S, Edachery S, Patel A, Kumar A, Shetty P, Santra MK. (2021) FBXL20 promotes breast cancer malignancy by inhibiting apoptosis through degradation of PUMA and BAX. J Biol Chem. 297, 101253. |
|
6. Islam, S., Dutta, P., Chopra, K., Sahay, O., Rapole, S., Chauhan, R. and Santra MK. (2021) Co-operative binding of SKP1, Cullin1 and Cullin7 to FBXW8 results in Cullin1-SKP1-FBXW8-Cullin7 functional complex formation that monitors cellular function of β-TrCP1. Int J Biol Macromol. 190, 233-243. |
|
7. Islam, S., Dutta, P., Copara, K., Rapole, S., Chauhan, R., and Santra M. K. (2021) FBXW8 regulates G1 and S phases of cell cycle progression by restricting β-TrCP1 function. FEBS J. 288, 5474 – 5497. |
|
8. Paul, D., Rapole, S. and Santra M. K. (2020) A new player in an old story: FBXO16 prevents breast cancer tumorigenesis through disrupting cellular function of nuclear β-Catenin. J. Expt. Pathology, 1, 22 – 27. |
|
9. Dutta, P., Islam, S., Choppara, S., Sengupta, P., Kumar, A., Kumar, A., Wani, M. R., Chatterjee, S. and Santra, M. K. (2019) The tumor suppressor FBXO31 preserves genomic integrity by regulating DNA replication and segregation through precise control of cyclin A levels. J. Biol. Chem. 294, 14879–14895. |
|
10. Paul D., Islam, S., Manne, R. K., Dinesh, U. S., Malonia, S. K., Maity, B., Boppana, R., Raople, S., Shetty, P. and Santra, M. K. (2019) F-box protein FBXO16 functions as a tumor suppressor by attenuating nuclear β-catenin function. J. of Pathology, 284, 266–279. |
|
11. Paul, D., Bargale, A. B., Raople, S., Shetty, P. and Santra, M. K. (2019) Protein phosphatase 1 regulatory subunit SDS22 inhibits breast cancer cell tumorigenesis by functioning as a negative regulator of the AKT signaling pathway. Neoplasia, 21, 30 – 40. |
|
12. Choppara, S., Malonia, S. K., Sankaran, G., Green M. R. and Santra M. K. (2018) Degradation of FBXO31 by APC/C is regulated by AKT- and ATM-mediated phosphorylation. Proc. Natl. Acad. Sci. USA., 115, 998 – 1003. |
|
13. Choppara, S., Sankaran, G., Manne, R., Dutta, P., Singh, S. and Santra M. K. (2018). The SCFFBXO46 ubiquitin ligase complex mediates degradation of the tumor suppressor FBXO31 and thereby prevents premature cellular senescence. J. Biol. Chem.293, 16291-16306. |
|
14. Meshram, S. N., Paul, D., Manne, R., Choppara, S., Sankaran, G., Agrawal, Y. and Santra, M. K. (2017). FBXO32 activates NF-κB through IκBα degradation in inflammatory and genotoxic stress. Int. J. Biochem. Cell Biol., 92, 134 – 140. |
|
15. Paul, D., Ghorai, S., Dinesh, U. S., Shetty, P., Chattopadhyay, S. and Santra, M. K. (2017) Cdc20 directs proteasome-mediated degradation of the tumor suppressor SMAR1 in higher grades of cancer through the anaphase promoting complex. Cell Death Dis. 8, e2882. |
|
16. Manne, R. K., Agrawal, Y., Bargale, A., Patel, A., Paul, D., Gupta, N. A., Rapole, S., Seshadri, V., Subramanyam, D., Shetty, P., and Santra, M. K. (2017) A MicroRNA/Ubiquitin Ligase Feedback Loop Regulates Slug-Mediated Invasion in Breast Cancer. Neoplasia, 19, 483 – 495. |
|
17. *Santra, M. K., Banerjee, A., Rahaman, O. and Panda, D. (2005) Unfolding pathways of human serum albumin: Evidence for sequential unfolding and folding of its three domains. Int. J. Biol. Macromol. 37, 200 – 204. |
|
18. Malonia, S. K., Dutta, P., Santra, M. K.* and Green, M. R*. (2015) F-box protein FBXO31 directs degradation of MDM2 to facilitate p53-mediated growth arrest following genotoxic stress, Proc. Natl. Acad. Sci. USA, 112, 8632 – 8637. |
|
19. Santra, M. K., Wajapeyee, N. and Green, M. R. (2009) F-Box protein FBXO31 mediates cyclin D1 degradation to induce G1 arrest following DNA damage, Nature, 459, 722-725. |
|
20. Santra, M, K. and Panda, D. (2007) Acid-induced loss of functional properties of bacterial cell division protein FtsZ: evidence for an alternative conformation at acidic pH. Proteins 67, 177 – 188. |
|
21. Santra, M. K., Banerjee, A., Krishnakumar, S. S., Rahaman, O. and Panda, D. (2004) Multiple-probe analysis of folding and unfolding pathways of human serum albumin. Evidence for a framework mechanism of folding. Eur J. Biochem. 271, 1789 – 1797. |
|
22. Santra, M. K., Beuria, T. K., Banerjee, A. and Panda, D. (2004) Ruthenium red-induced bundling of bacterial cell division protein, FtsZ. J. Biol. Chem. 279, 25959 – 25965. |
|
23. Santra, M. K. and Panda, D. (2003) Detection of an intermediate during unfolding of bacterial cell division protein FtsZ: loss of functional properties precedes the global unfolding of FtsZ. J. Biol. Chem. 278, 21336 – 21243. |
| 24. Santra, M. K., Dasgupta, D. and Panda, D. (2006) Pyrene Excimer Fluorescence of Yeast Alcohol Dehydrogenase: A Sensitive Probe to Investigate Ligand Binding and Unfolding Pathway of the Enzyme. Photochem. Photobiol. 82, 480 – 486.
|
|
25. Santra, M. K., Dasgupta, D. and Panda, D. (2005) Deuterium oxide promotes assembly and bundling of FtsZ protofilaments. Proteins 61, 1101-1110. |
|
26. Sivakrishna, B., Islam, S., Santra, M. K.* and Pal, S.* (2019) Synthesis and cytotoxic evaluation of apioarabinofuranosyl pyrimidines. Drug Dev. Res. 81, 274–282. |
|
27. Sivakrishna, B. Islam, S., Panda, A., Saranya, M., Santra, M. K.* and Pal. S* (2018) Synthesis and Anticancer Properties of Novel Truncated Carbocyclic Nucleoside Analogues. Anticancer Agents Med Chem. 18, 1 – 7. |
|
28. Gorai, S., Paul, D., Borah, R., Haloi, N., Santra, M. K.* and Manna, D.* (2018) Role of Cationic Groove and Hydrophobic Residues in Phosphatidylinositol-Dependent Membrane-Binding Properties of Tks5-Phox Homology Domain. Chemistry Select, 3, 1205 – 1214. |
|
29. Valkute, T. R. Eswar K. Aratikatla, E. K. Gupta, N. A. Ganga, S. Santra, M. K.* and Bhattacharya, A. K.* (2018) Synthesis and anticancer studies of Michael adducts and Heck arylation products of sesquiterpene lactones, zaluzanin D and zaluzanin C from Vernonia arborea. RSC Adv., 8, 38289 |
|
30. Ghosh, C., Gupta, N., Mallick, A., Santra, M. K.* and Basu, S.* (2018) Self-Assembled Glycosylated Chalcone–Boronic Acid Nanodrug Exhibits Anticancer Activity through Mitochondrial Impairment ACS Appl. Bio Mater., 1, 347 – 355. |
|
31. Paul, D., Chanukuppa, V., Reddy, P. J., Taunk, K., Adhav, R., Srivastava, S., Santra, M. K.* and Rapole, S.* (2016) Global proteomic profiling identifies etoposide chemoresistance markers in non-small cell lung carcinoma, J. Proteomics, 138, 95-105. |
|
32. Ghosh, C., Gupta, N., More, P., Sengupta, P., Mallick, A., Santra, M. K.* and Basu, S.* (2016) Engineering and In Vitro Evaluation of Acid Labile Cholesterol Tethered MG132 Nanoparticle for Targeting Ubiquitin‐Proteasome System in Cancer. Chemistry Select, 1, 5099-5106. |
|
33. Gorai, S., Paul, S., Sankaran, G., Borah, R., Santra, M. K.* and Manna, D.* (2015) Inhibition of phosphatidylinositol-3,4,5-trisphosphate binding to the AKT pleckstrin homology domain by 4-amino-1,2,5-oxadiazole derivatives. Med. Chem. Commun. 6, 1798 – 1808. |
|
34. Panda, A., Islam, S., Santra, M. K.* and Pal, S.* (2015) Lead tetraacetate mediated one pot oxidative cleavage and acetylation reaction: an approach to apio and homologated apio pyrimidine nucleosides and their anticancer activity. RSC Adv., 5, 82450 – 82459. |
|
35. Paul D, Kumar A, Gajbhiye A, Santra, M. K.* and Srikanth, R.* (2013) Mass spectrometry-based proteomics in molecular diagnostics: discovery of cancer biomarkers using tissue culture. Biomed Res Int. 2013:783131. |
|
36. Gorai, S., Paul, D., Haloi, N., Borah, R., Santra, M. K.* and Manna, D*. (2016) Mechanistic insights into the phosphatidylinositol binding properties of the pleckstrin homology domain of lamellipodin, Molecular BioSystems, 112, 8632 – 8637. 37. T |
|
37. Tripathy, S. K., Surada, R. K., Manne, R. K, Mobin, S. M., Santra, M. K*. and Patra, S*. (2013) Synthesis, characterisation and biological activities of [(p-cym)RuX(μpz4lut)]n+ and[{(p-cym)RuX}2(μ-pz4lut)]n+ (X = Cl, H2O and pz4lut =α,α,α',α'-tetra(pyrazol-1-yl)-2,6-lutidine). Dalton Trans, 42, 14081-14091. |
|
38. Kumari, A., Shriwas, O., Sisodiya, S., Santra, M. K., Guchhait, S. K., Dash, R., Panda, D. (2021) Microtubule-targeting agents impair kinesin-2-dependent nuclear transport of β-catenin: Evidence of inhibition of Wnt/β-catenin signaling as an important antitumor mechanism of microtubule-targeting agents. FASEB J. 35, e21539. |
|
39. Chanukuppa, V., Taware, R., Taunk, K., Chatterjee, T., Sharma, S., Somasundaram, V., Rashid, F., Malakar, D., Santra, M. K. and Rapole S. Proteomic Alterations in Multiple Myeloma: A Comprehensive Study Using Bone Marrow Interstitial Fluid and Serum Samples. Front Oncol. 2021 Jan 29;10:566804. doi: 10.3389
|
|
40. Waghela, B. N., Foram U Vaidya, F. U., Agrawal, Y., Santra, M. K., Mishra, V. and Pathak, C. (2020) Molecular insights of NADPH oxidases and its pathological consequences. Cell Biochem. Funct. 39, 218–234. |
|
41. Akhtar N, Pradhan N, Barik GK, Chatterjee S, Ghosh S, Saha A, Satpati P, Bhattacharyya A, Santra MK, Manna, D. (2020) Quinine-Based Semisynthetic Ion Transporters with Potential Antiproliferative Activities. ACS Appl. Mater Interfaces 12, 25521-25533. |
|
42. Sivakrishna, B., Shukla, M., Santra, M. K. and Pal, S. (2020) Design, synthesis and cytotoxic evaluation of truncated 3'-deoxy- 3', 3' difluororibofuranosyl pyrimidine nucleosides Carbohydr Res. 2020 Aug 16;497:108113. doi: 10.1016/j.carres.2020.108113. |
|
43. Chanukuppa, V., Paul, D., Taunk, K., Chatterjee, T., Sharma, S., Shirolkar, A., Islam, S., Santra, M. K. and Rapole, S. (2020) Proteomics and functional study reveal marginal zone B and B1 cell specific protein as a candidate marker of multiple myeloma. Int. J. Oncol. 57, 325–337. |
|
44. Kotammagari, T. K., Paul, S., Barik, G. K., Santra, M. K., Bhattacharya, A. K. (2020) Synthesis of artemisinic acid derived glycoconjugates and their anticancer studies. Org. Biomol. Chem. 18, 2252–2263. |
|
45. Rohini, V., Gujar, V., Pathan, H., Islam, S., Tawre, M., Pardesi, K., Santra, M. K. and Ottoor, D. (2019) Bioimaging Applications of Carbon dots (C. dots) and its Cystamine Functionalization for the Sensitive Detection of Cr(VI) in Aqueous Samples. J. Fluoresc. 29, 1381-1392. |
|
46. Chanukuppa, V., Paul, D., Taunk, K., Chatterjee, T., Sharma, S., Kumar, S., Santra, M. K. and Rapole, S. (2019) XPO1 is a critical player for bortezomib resistance in multiple myeloma: A quantitative proteomic approach. J Proteomics. 209, 103504. |
|
47. Kumari, A., Srivastava, S., Manne, R. K., Sisodiya, S., Santra, M. K., Guchhait, S. K. and Panda D. (2019) C12, a combretastatin-A4 analog, exerts anticancer activity by targeting microtubules. Biochem. Pharmacol. 170, 113663. |
|
48. Majumdar, D., Agrawal, Y., Thomas, R., Ullah, Z., Santra, M. K., Das, S., Pal, T. K., Bankura, K. and Mishra, D. (2019) Syntheses, characterizations, crystal structures, DFT/TD‐DFT, luminescence behaviors and cytotoxic effect of bicompartmental Zn (II)‐dicyanamide Schiff base coordination polymers: An approach to apoptosis, autophagy and necrosis type classical cell death. Appl. Organometal. Chem., 34, e5269. /doi.org/10.1002/aoc.5269 |
|
49. Shinde, P., Melinkeri, S., Santra M. K., Kale, V. and Limaye L. (2019) Autologous Hematopoietic Stem Cells Are a Preferred Source to Generate Dendritic Cells for Immunotherapy in Multiple Myeloma Patients. Front Immunol. 10, 1079. |
|
50. Manna, S. C., Mistri, S., Patra, A., Mahish, M. K., Saren, D., Manne, R. K., Santra, M. K., Zangrando, E. and Puschmann, H. (2019) Synthesis, structure, DNA/protein binding, molecular docking and in vitro anticancer activity of two Schiff base coordinated copper(II) complexes. Polyhedron 171, 77–85. |
|
51. Bhunia, A., Mistri, S., Manne, R. K. Santra, M. K. Manna, S. C. (2019) Synthesis, crystal structure, cytotoxicity study, DNA/protein binding and molecular docking of dinuclear copper(II) complexes. Inorganica Chimica Acta, 491, 25–33. |
|
52. Roshni V, Misra, S., Santra, M. K. and Ottoor, V. (2019) One pot green synthesis of C-dots from groundnuts and its application as Cr (VI) sensor and in vitro bioimaging agent. J. of Photochem. & Photobiol. A: Chemistry, 373, 28–36. |
|
53. Chanukuppa, V., Taware, R., Chatterjee, T., Sharma, S., More, T. H., Taunk, K., Kumar, S., Santra, M. K. and Rapole, S. (2018) Current understanding of the potential of proteomics and metabolomics approaches in cancer chemoresistance: A focus on Multiple Myeloma. Curr Top Med Chem. 18, 2584 - 2598. |
|
54. Taye, N. Alam, A., Ghorai S, Chatterji DG, Parulekar A, Mogare D, Singh S, Sengupta P, Chatterjee S, Bhat MK, Santra, M. K. Salunkhe PB, Finston SK and Chattopadhyay, S. (2018) SMAR1 inhibits Wnt/β-catenin signaling and prevents colorectal cancer progression. Oncotarget, 9, 21322-21336. |
|
55. Mistri, S., Patra, A., Santra, M. K., Paul, D., Zangrando, E., Puschmann, H. and Manna, S. C. (2018). DNA/Protein Binding, Molecular Docking and Cytotoxicity Studies of Piperazinyl-Moiety-Based Copper(II) Complexes. Chemistry Select, 3, 9102 – 9112. |
|
56. Jhulki, L., Dutta, P. Santra, M. K., Cardoso, M. H., Oshiro, G. N., Franco, O. L., Bertolasi, V., Isab, A. A., Bielawski, C. W. and Dinda, J. (2018) Synthesis and cytotoxic characteristics displayed by a series of Ag(I)-, Au(I)- and Au(III)-complexes supported by a common N-heterocyclic carbine. New Journal of Chemistry, 42, 13948-13956. |
|
57. Das, U., Pattanayak, P., Santra, M. K. and Chattopadhyay, S. (2018) Synthesis of new oxido-vanadium complexes: catalytic properties and cytotoxicity. J. Chemical Res., 42, 57 – 62. |
|
58. Baravkar, S. B., Wagh, M. A., Paul, D., Santra M. K. and Sanjayan, G. J. (2018) Synthesis and anticancer activity of conformationally constrained Smac mimetics containing pseudo b turns. Tetrahedron Letters, 59, 3473–3476. |
|
59. More, T. H., RoyChoudhury, S., Christie, J., Taunk, K., Mane, A., Santra, M. K., Chaudhury, K. and Rapole, S. (2017) Metabolomic alterations in invasive ductal carcinoma of breast: A comprehensive metabolomic study using tissue and serum samples. Oncotarget, 9, 2678-2696. |
|
60. Gajbhiye, A., Dabhi, R., Taunk, K., Jagadeeshaprasad, M. G., RoyChoudhury, S., Mane, A., Bayatigeri, S., Chaudhury, K., Santra, M. K. and Rapole, S. (2017) Multipronged quantitative proteomics reveals serum proteome alterations in breast cancer intrinsic subtypes. J. Proteomics, 163, 1 – 13. |
|
61. Rai, A., Kapoor, S., Naaz, A., Santra, M. K. and Panda, D. (2017) Enhanced stability of microtubules contributes in the development of colchicine resistance in MCF-7 cells. Biochem Pharmacol., 132, 38 – 47. |
|
62. Augustine, T., Chaudhary, P., Gupta, K., Islam, S., Ghosh, P., Santra, M. K. and Mitra, D. (2017) Cyclin F/FBXO1 interacts with HIV-1 Vif and restricts progeny virion infectivity by ubiquitination and proteasomal degradation of Vif through SCF (Cyclin F) E3 ligase machinery. J. Biol. Chem., 292, 5349 – 5363. |
|
63. More, T.H., Bagadi, M., RoyChoudhury, S., Dutta, M., Uppal, A., Mane, A., Santra, M.K., Chaudhury, K. and Rapole, S. (2017) Comprehensive quantitative lipidomic approach to investigate serum phospholipid alterations in breast cancer. Metabolomics, 13, 3 – 9. |
|
64. Gajbhiye, A., Dabhi, R., Taunk, K., Vannuruswamy, G., RoyChoudhury, S., Adhav, R., Seal, S., Mane, A., Bayatigeri, S., Santra, M. K., Chaudhury, K. and Rapole, S. (2016) Urinary proteome alterations in HER2 enriched breast cancer revealed by multipronged quantitative proteomics. Proteomics 16, 2403-2418. |
|
65. Nusrat S, Zaidi N, Zaman M, Islam S, Ajmal MR, Siddiqi MK, Santra MK, Khan RH. (2016) Repositioning nordihydroguaiaretic acid as a potent inhibitor of systemic amyloidosis and associated cellular toxicity. Arch Biochem Biophys. 612, 78 – 90. |
|
66. Pendharkar, N., Gajbhiye, A., Taunk, K., RoyChoudhury, S., Dhali, S., Seal, S., Mane, A., Abhang, S, Santra, M. K., Chaudhury, K. and Rapole, S. (2016) Quantitative tissue proteomic investigation of invasive ductal carcinoma of breast with luminal B HER2 positive and HER2 enriched subtypes towards potential diagnostic and therapeutic biomarkers, Journal of Proteomics, 132, 112-130. |
|
67. Mistri, S., Paul, A., Bhunia, A., Manne, R. K., Santra, M. K., Puschmann, H. and Manna, S. C. (2016) A combined experimental and theoretical investigation on the Cu (II) sensing behavior of a piperazinyl moiety based ligand, and catecholase and biological activities of its Cu (II) complex in combination with pyridine 2, 5-dicarboxylate, Polyhedron, 104, 63 – 72. |
|
68. Nakka, K. K., Chaudhary, N., Joshi, S., Singh, J., Chatterjee, S., Malhotra, R., De, A., Santra, M. K., Dilworth, F. J. and Chattopadhyay, S. (2015) Nuclear matrix-associated protein SMAR1 regulates alternative splicing via HDAC6-mediated deacetylation of Sam68 Proc. Natl. Acad. Sci. USA, 112, E3374 – E3383. |
|
69. Bhunia, A., Manna, S., Mistri, S., Paul, A., Manne, R. K., Santra, M. K., Bertolasi, V. and Manna, S. C. (2015) Synthesis, characterization, TDDFT calculation and biological activity of tetradentate ligand based square pyramidal Cu(II) complexes. RSC Adv., 5, 67727 – 67737. |
|
70. Gorai, S., Bagdi, P. R., Borah, R., Paul, D., Santra, M. K., Khan, A. T. Manna, D. (2015) Insights into the inhibitory mechanism of triazole-based small molecules on phosphatidylinositol-4,5-bisphosphate binding pleckstrin homology domain. Biochemistry and Biophysics Reports, 2, 75–86. |
|
71. Kale, S. S., Jedhe, G. S., Meshram S. N., Santra, M. K., Hamel, E. and Sanjayan, G. J. (2015) Novel hybrid nocodazole analogues as tubulin polymerization inhibitors and their antiproliferative activity. Bioorg. Med. Chem. Lett. 25, 1982 – 1985. |
|
72. Dinda, J., Adhikary, D. S. Roymahapatra, G., Nakka, K.K., and Santra, M. K.(2014) Synthesis, Structure, Electrochemistry and Cytotoxicity Studies of Ru(II) and Pt(II)-N- Heterocyclic Carbene Complexes of CNC-Pincer Ligand. Inorganica Chimica Acta, 413, 23 – 31. |
|
73. Jedhe, G.S., Paul, D., Gonnade, R. G., Santra, M. K., Hamel, E., Nguyen, T. L., and Sanjayan, G. J. (2013) Correlation of hydrogen-bonding propensity and anticancer profile of tetrazole-tethered combretastatin analogues. Bioorg Med. Chem. Lett., 23, 4680-4684. |
|
74. Kumar, P., Gorai, S., Santra, M. K., Mondal, B. and Manna, D. (2012) DNA binding, nuclease activity and cytotoxicity studies of Cu(II) complexes of tridentate ligands, Dalton Transactions, 41, 7573-7581. |
|
75. Palakurty, R., Wajapeyee, N., Santra, M. K., Ling, L., Gazin, C, Gobeil, S. and Green, M. R. (2009) Epigenetic silencing of RASSF1A tumor suppressor gene through HOXB3-mediated induction of DNMT3b expression. Molecular Cell 36, 219-230. |
|
76. Welch, C., Santra, M. K., Zhu, X., El-Assaad, W., Huber, W. E., Keys, R. A., Jose, G. Teodoro, J. G. and Green, M. R. (2009) Identification of a protein, G0S2, that lacks Bcl-2 homology domains and interacts with and antagonizes Bcl-2. Cancer Research, 69, 6782-6789. |
|
77. Hart, D. Santra, M. K. and Green, M. R. (2009) Selective interaction of TAF3 and TRF3 regulates the embryonic development. Developmental Dynamics, 238, 2540-2549. |
|
78. Beuria, T. K., Shah, J. H., Santra, M. K., Kumar, V. and Panda, D. (2006) Effects of pH and ionic on the assembly and bundling of FtsZ protofilaments: a possible role of electrostatic interactions in the bundling of protofilaments. Int. J. Biol. Macromol. 40, 30 – 39. |
|
79. Panda, D., Rathinasamy, K., Santra, M. K. and Wilson, L. (2005) Kinetic suppression of microtubule dynamic instability by griseofulvin: implications for its possible use in the treatment of cancer. Proc. Natl. Acad. Sci. USA 102, 9878 – 9883. |
|
80. Beuria, T. K., Santra, M. K. and Panda, D. (2005) Sanguinarine Blocks Cytokinesis in Bacteria by Inhibiting FtsZ Assembly and Bundling. Biochemistry 44, 16584 – 16593. |
|
81. Mukherjee, A., Santra, M. K., Beuria, T. K. and Panda, D. (2005) A natural osmolyte trimethylamine N-oxide promotes assembly and bundling of the bacterial cell division protein, FtsZ and counteracts the denaturing effects of urea. FEBS J. 272, 2760 – 2772. |
Book Chapters
| 1. Kalita, B. Barik, GK. Sharma, T. Taunk, T. Santra, M. K. Rapole, S. (2022) Systemic metabolomic changes associated with chemotherapy: Role in personalized therapy, Wiley |
| 2. Samanta, S. Mahata, R and Santra, M. K. (2022) Metabolism and Epigenetic Regulation: Implications in Cancer, Subcellular Biochemistry book series, Publisher – Springer |
| 3. Agrawal, Y and Santra, M. K. (2019) Interplay of Ubiquitination and SUMOylation with miRNAs, Publisher -Caister Academic Press, Pages: 217-230 |
Awards/Honours
2021: Fellow of National Academy of Science (FNASc), Allahabad, 2021
2019: National Bioscience Award for Career Development, Department of Biotechnology, Government of India.
2010: Charles A. King Trust Postdoctoral Research Fellowship Award, USA
2010: Ramalingaswami Fellowship, Department of Biotechnology, Government of India.
2009: Innovative Young Biologist Award for the year 2008 from Department of Biotechnology, Government of India.
2006: Young Scientist Award in New Biology from Indian Science Congress Association.
2006: Best research scholar award from IIT Bombay for the year 2006
2005: IUBMB-Sigma Aldrich Corporation Travel Scholarship for attending FEBS Forum for Young Scientists and 30th FEBS Congress-9th IUBMB Conference which was held at
Budapest, Hungary, June 2005.
2001: Junior research fellowship from CSIR to peruse Ph.D. from IIT Bombay, India
1999: MHRD fellowship (GATE, All India rank 4th) to peruse M. Tech. from IIT Bombay, India.
Memberships
- Life member for the Society of Biological Chemists
- Life member for Indian Association for Cancer Research
- Life member for Indian Society of Cell Biology
- Life member for the Indian Science Congress Association
- Life member of Indian Society of Translational Research (ISTR)
- Life member of Proteomic Society of India (PSI)
- Life member of the society for Free Radical Research-India (SFRR-India)
Lab Members

Nikhil Baban Ghate, MK Bhan Young Researcher Fellow (Post-Doc Fellow) , DOJ- 1st April 2022, PhD Biochemistry (2016), University of Calcutta (WB).
Ganesh Kumar Barik, UGC-SRF, DOJ- 1st August 2016, M.Sc. Zoology (2016), Banaras Hindu University (UP).
Tanisha Sharma, UGC-SRF, DOJ-1st March 2018, M.Sc. Life Sciences (2016), Jawaharlal Nehru University (ND).
Snigdha Samanta, CSIR-JRF, DOJ-2nd March 2020, M.Sc. Biotechnology (2019), Jawaharlal Nehru University (ND).
Rumpa Mahata, CSIR-JRF, DOJ- 12th October 2020, M.Sc. Biotechnology (2019), Visva-Bharati (WB).
Brindaban Das, DBT-JRF, DOJ-1st March 2022, M.Sc. Zoology (2020), Visva-Bharati (WB).
Project JRF
Chandra Biswas, DOJ-27th December 2021, M.Sc. Zoology (2021), Cotton University, Assam.
Lab Alumni
Alumni PhD Fellows
Debasish Paul, UGC-SRF, DOJ- 5th March 2012, M.Sc. Plant Mol. Bio. & Biotechnology (2011), Bose Institute and Univ. Of Calcutta, (WB)
Neha Anilkumar Gupta, UGC-JRF, DOJ-17th July 2013, M.Sc. Biochemistry (2011), University of Hyderabad (AP)
Parul Dutta, UGC-SRF, DOJ- 2nd Nov. 2011, M.Sc Zoology (2010), Banaras Hindu University (UP)
Rajesh kumar Manne, UGC-SRF, DOJ-1st Jan.2011, M.Sc Biochemistry & Molecular, Biology (2010), Pondicherry University, Pondicherry
Sachin N. M. Meshram, CSIR-SRF, DOJ- 3rd Nov. 2011, M.Sc Biotechnology (2011), Sant Gadge Baba Amravati University (MH)
Sehbanul Islam, DBT-JRF, DOJ- 7th March 2014, M.Sc. Biotechnology (2013) Aligarh Muslim University, Aligarh (UP)
Srinadh Choppara, CSIR-SRF, DOJ- 1st Nov. 2011, M.Sc Biotechnology (2010) JNTU Hyderabad (AP)
Yashika Agarwal, DBT-JRF, DOJ- 2014, M.Sc Biotechnology, (2012) Amity University, Lucknow (UP)
Alumni Post-Doctoral Fellows
Bhargav Avinash Kumar (2011)
Jyotika Rajawat (2016)
Mahua Rani Das (2017)
Shweta Misra (2017)
Project JRF
Asha Patel, DOJ- 26th April 2012, MSc Biotechnology, (2010), Dr. Hari Singh Gour University, Sagar, (MP)
Sushrita Roy Muhury, DOJ- 7th September 2018, M.Sc Biotechnology (2018), St. Xavier’s College (WB)
Kaustubh Sanjay Nadkarni, DOJ-28th October 2018, M.Sc Biotechnology (2016), Pune University, Maharashtra.
Project Assistant
S. Ganga, DOJ- 23rd Oct. 2013, B.Tech Biotechnology (2013), Bharathidasan University, Trichy (TN)
Students trained in our Lab
Six month trainees
Bhargav Waghela (2011-12)
Lakshmana Kumar Kasarapu (2011-12)
Padmaja Murali (2013)
Shringika Mishra (2014)
Awadesh Pratap Sing (2014)
Sharad Shriram Tat (2014)
Sayantani Bhowmik (2014)
Rishita Chatterjee (2014)
Sonali Kale (2015)
Kavita Mishra (2016)
Vidusha Bhaduria (2016)
Gopalakrishnan (2016)
Shreya Panicker (2017)
Shrutika Kiran Sasane (2017)
Vandana Jhalora (2017)
Vharte Bharat Kisan (2018)
Manisha Joshi (2018)
Payal Shaw (2018)
Rashida Makda (2018)
Sanjana Shajan (2018)
Apurva Bhat (2019)
Gaurishankar Bhaskar (2019)
Dhiraj Sanjay Powar (2019)
Sakshi Bubna (2020)
Amirtha Varshni P T R (2021)
Chandra Biswas (2021)
Ruchi Verma (2021)
IAS Research Fellows
Sreeparna Podder (2015)
Kavita Mishra (2016)
Glancis Luzeena (2017)
G. Koushik (2018)
B. Madhumitha (2018)
Joshuah Fialho (2019)
Aparna Banerjee (2020)
Chandra Biswas (2020)
Varsha N (2021)
Back
Last updated On : 04 July 2022 08:47


